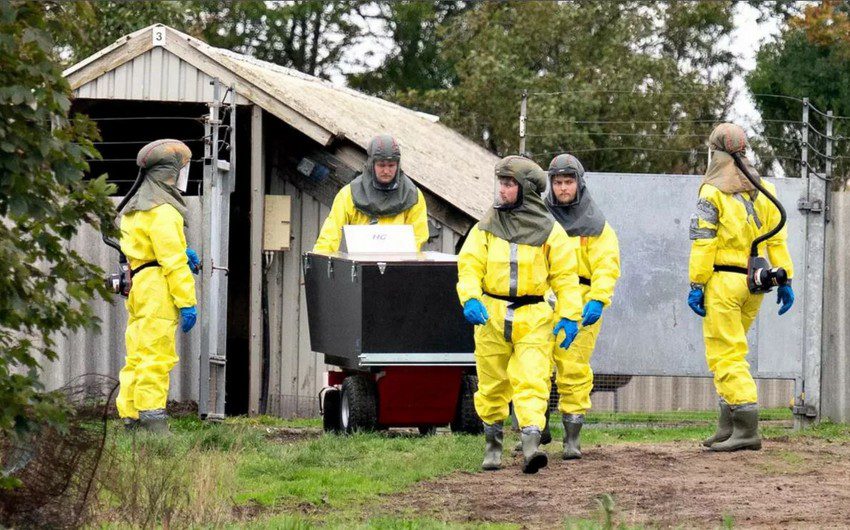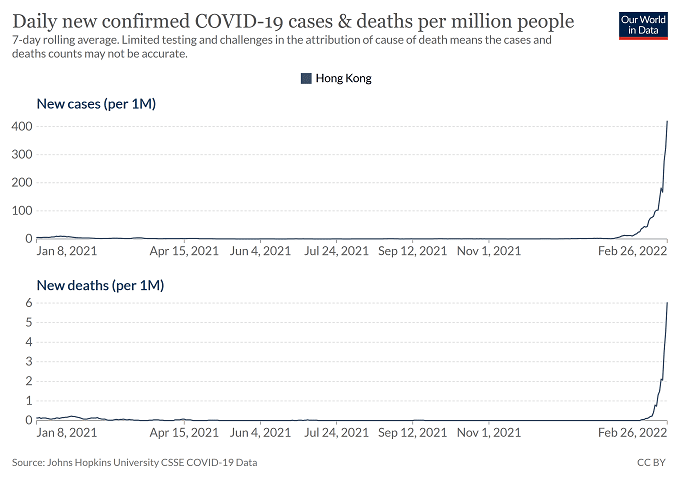
France: B.1.160 outbreak may have begun in mink
“based on genomic, geographical and temporal coincidences, that the source of the Marseille-4 (B.1.160) epidemic was the mink”.
The Marseille-4 variant (B.1.160) was the most prevalent SARS-CoV-2 variant in our geographical area in 2020 and was only outdone by the Alpha (UK, B.1.1.7) variant in 2021. It replaced the Marseille-1 variant in August 2020 and faded away in April 2021. It was first detected in south-west France and in Marseille.
Its emergence coincided with the re-increase in France of SARS-CoV-2 incidence starting with a department in northwest France (Mayenne) close to the Eure-et-Loir department where a mink farm was found to be massively infected with SARS-CoV-2. The only genome from this farm, obtained from a mink sampled in mid-November 2020 but released in March 2021, is that of a Marseille-4 variant (GISAID Accession no. EPI_ISL_1392906).
In addition, the co-occurrence of 13 hallmark mutations in the Marseille-4 variant without any available genome showing a gradual appearance of these mutations suggested that genetic evolution had been overlooked. These data led us to conclude, based on genomic, geographical and temporal coincidences, that the source of the Marseille-4 epidemic was the mink.
The Marseille-4 variant was progressively replaced from January 2021 by the epidemic linked to the Alpha (UK, B.1.1.7) variant. Meanwhile, it circulated in 76 countries and accounted for 31,299 genomes in the GISAID database as of 30 June 2021, mostly obtained from European countries [in 29,906 cases].
Preprint: Analysis of SARS-CoV-2 Variants From 24,181 Patients Exemplifies the Role of Globalization and Zoonosis in Pandemics
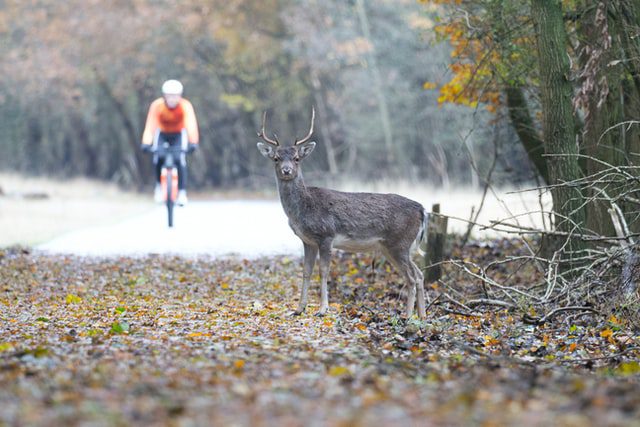
Interestingly, the closest relatives to the ON deer were human and mink-derived samples from nearby Michigan back in 2020.
We also identified a single human case that was very similar to our deer samples and came from the same time-frame and region as the deer samples /4 pic.twitter.com/lJt52UNHvy
— Finlay Maguire (@FinlayM) February 26, 2022
Preprint: Highly divergent lineage of SARS-CoV-2 in deer with potential deer-to-human transmission
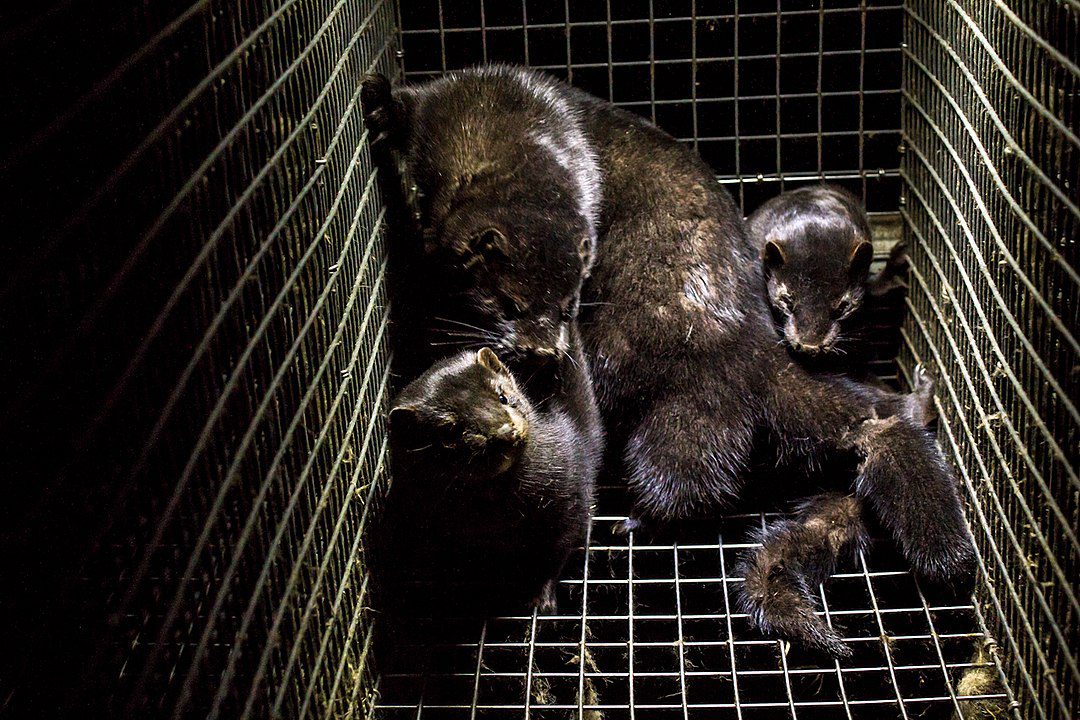
Image by Otwarte Klatki, CC BY 2.0, via Wikimedia Commons