China: 50 SARS-CoV-2 sequences uploaded to Gisaid
Gisaid have published a report recently on fifty SARS-CoV-2 genome sequences uploaded by China which seems to have gone largely unnoticed by the mainstream media. The report sheds a bit more light on what might be happening in China, not just in Shanghai, but across the entire country.
The report outlines some of the SARS-CoV-2 variants the Chinese are currently tackling, and there are certainly some surprises in the list.
“Six genomes were assigned as Delta (from four distinct lineages), and 44 genomes were assigned as Omicron (from seven distinct lineages). No highly divergent new variants were found.”
That’s probably a higher percentage of Delta sequences than many might have imagined, and mirrors the situation in Great Britain early in the new year of 2022, after which a number of Omicron & Delta (Deltacron) recombinants were discovered.
The creation of Deltacron variants through recombination could become a problem with this mix of lineages circulating. That may be one of the reasons why the Chinese CDC published a paper on this very subject recently, which we covered in this post: China CDC: Deltacron is a “Grey Rhino” event.
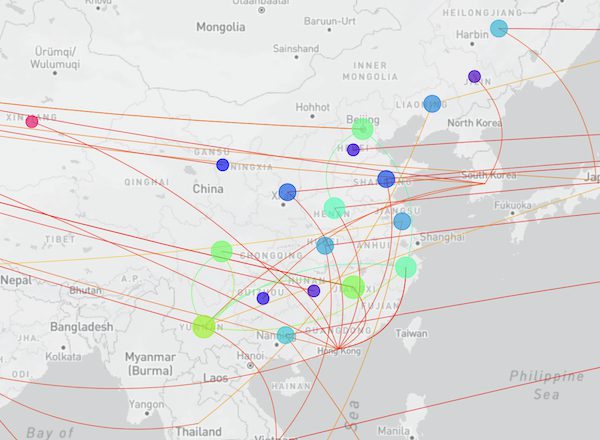
“Searching the 50 new hCoV-19 genome sequences with GISAID’s AudacityInstant against the 10.2 million genomes in the EpiCoV database found a total of 14,176 closely related genomes. The graphic below shows the distribution of countries of the closest hits of which the dark blue bars correspond to identical sequence hits from abroad and possible transmission links”:
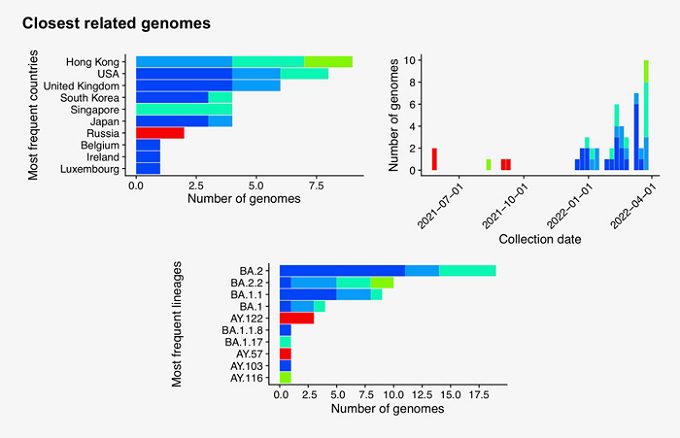
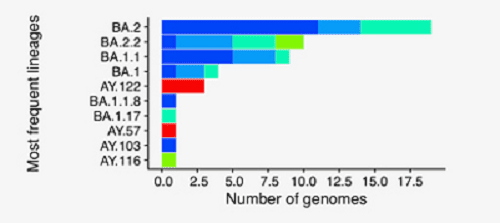
The two most common lineages in the sequences are BA.2 and BA.2.2, and it is perhaps the latter of the two that could potentially be the more troubling, because BA.2.2 is the lineage that caused so many infections and deaths amongst the elderly population of Hong Kong recently.
We covered BA.2.2 in the following posts:
Hong Kong: Omicron BA.2 with S:I1221T mutation at 100% of recent sequences *6 UPDATES*
Hong Kong: Omicron BA.2 with S:I1221T mutation designated BA.2.2
The full Gisaid report on the Chinese sequences can be found here
Our opinion: Uploading 50 SARS-CoV-2 sequences is a good start, but the scientific community could do with thousands more sequences from China to be able to predict where this virus is headed with any degree of accuracy. The virus is the only winner if genome sequences aren’t shared.