Category: Japan
Censored by Youtube: Dr John Campbell on 14yr old Japanese girl who died two days after third dose of Pfizer Covid vaccine
Youtube have once again overstepped their authority by censoring Covid-19 vaccine data that doesn’t fit the global narrative. More...
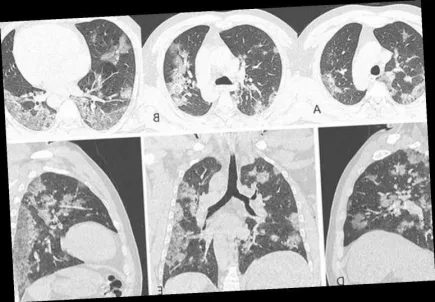
Sato Lab: Virological characteristics of XBB.1.5
Japan’s Sato Lab have announced new research into the Omicron XBB.1.5 ‘Kraken’ variant, confirming much about what is already known or suspected about its transmissibility. More...
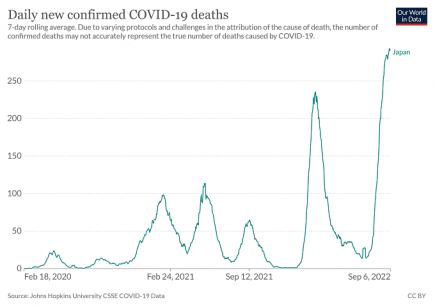
Record Covid deaths in Japan, pneumonia increases in Italy
Just two days ago, the WHO warned of an increase in Covid deaths across the globe. More...
Japan: 58% of COVID-19 deaths in under-20’s had no underlying conditions
A new report from Japan shows that the majority of Covid deaths in the under-20’s had no underlying medical conditions. More...
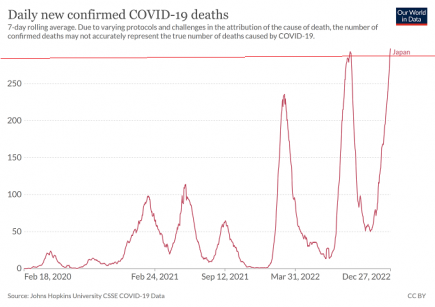
Japan: A new pandemic record for Covid-19 deaths *1 Update*
A new daily record for Covid-19 deaths has been set in Japan, and the link between deaths and cases appears to be breaking too. More...
Japan: new Covid wave begins with BF.5 as leading variant again
Japan is seeing a new wave of coronavirus rolling in, just weeks after the last devastating wave tailed off. More...
Japan: More than 800 infected with Covid after dance festival superspreader event
819 people were found to have been infected with coronavirus after a summer dance festival turned into a Covid superspreader event. More...
Japan: Half of Covid deaths in under 20’s had no underlying health problems *1 Update*
NHK are reporting that about half of under 20’s who died from Covid recently had no underlying health problems. More...
Japan: 89% of those who died of Covid in the latest wave had moderate symptoms
Japan says an increasing percentage of people are dying after developing moderate symptoms of COVID-19. More...
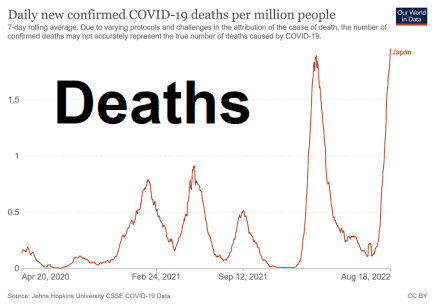
Japan: Record-breaking Covid-19 cases, hospitalizations and deaths
Japan has seen record-breaking Covid-19 cases, hospitalizations and deaths in the past few days. More...