Category: P.3
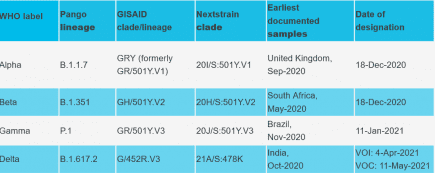
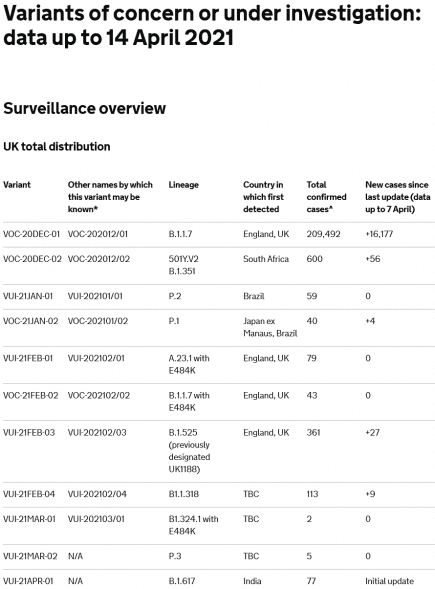
WHO announce new names for Sars-CoV-2 coronavirus variants
The WHO have announced new names for the confusingly named sars-cov-2 variants. More...
PHE: Two cases of Philippines coronavirus variant P.3 (PHL-B.1.1.28) found in England
“On 9 March, PHE noted a report of 33 cases of a new variant reported by the Philippines. More...
90 cases of Brazil related coronavirus variant PHL-B.1.1.28 in The Philippines
The Philippines now has all three variants of the coronavirus that have been fuelling record-breaking spikes in infections across the globe, and a Philippine variant (PHL-B.1.1.28) that has the same lineage as the infectious Brazilian variant. More...
Japan identifies a B.1.1.28 coronavirus strain with E484K and N501Y mutations from a traveller from the Philippines
The National Institute of Infectious Diseases, Japan, identified a B.1.1.28 strain with E484K and N501Y mutations from a SARS-CoV-2-positive sample collected on February 25, 2020 at a point of entry to Japan from a traveller from the Philippines(virus name : hCoV-19/Japan/IC-0824/2021, GISAID Accession ID: EPI_ISL_1198832) (1). More...
New coronavirus variant with E484K, N501Y & P681H mutations found in the Philippines, designated PHL-B.1.1.28
Here, we describe the emergence of a new SARS-CoV-2 lineage, mainly from the Central Visayas region of the Philippines. More...
New Philippines coronavirus variant has 484K and 501Y mutations
On Thursday, the Department of Health (DoH) in Central Visayas (The Philippines) confirmed that they have detected two mutations with “potential clinical significance” of SARS-CoV-2 that cause the disease. More...