Category: B.1.429
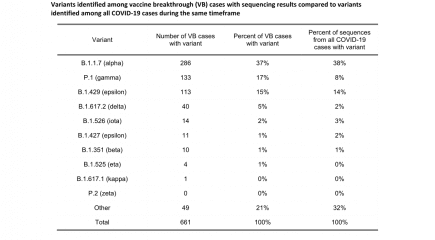
Vaccine breakthrough cases in Oregon by variant to 16th September 2021
Vaccine breakthrough data for Oregon state by Sars-CoV-2 variant up to 16th September 2021. More...
USA: 55% increase in Delta vaccine breakthrough cases in Washington State
This week’s update from Washington State showing vaccine breakthrough cases and which Sars-CoV-2 variant is causing them up to 11th August 2021. More...
USA: Washington State Vaccine Breakthroughs by Variant up to 28th July 2021
This week’s update from Washington State showing vaccine breakthrough cases and which Sars-CoV-2 variant is causing them up to 28th July 2021. More...
USA: Washington State Vaccine Breakthroughs by Variant up to 21st July 2021
This week’s update from Washington State showing vaccine breakthrough cases and which Sars-CoV-2 variant is causing them. More...
USA: Vaccine breakthough cases in Washington State by coronavirus variant to 30th June 2021
Washington State has recently published an update on the number of vaccine breakthrough cases. More...
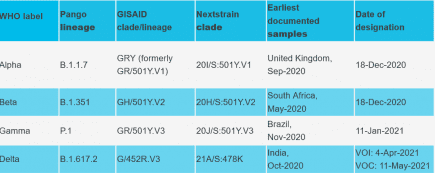
WHO announce new names for Sars-CoV-2 coronavirus variants
The WHO have announced new names for the confusingly named sars-cov-2 variants. More...
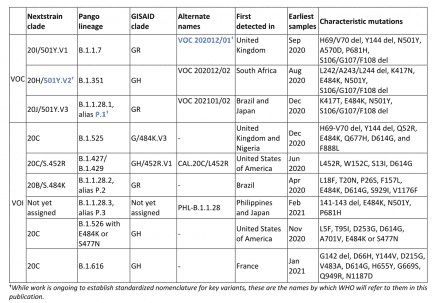
B1427/B1429 are listed as Variants of Concern by the CDC, as Variants of Interest (VOI) by WHO
The World Health Organization lists as B.1.427 and B.1.429 as Variants of Interest. More...
L452R mutation is present in more than 400 SARS-CoV-2 coronavirus genomes isolated from over 20 countries
Scientists have examined the Nextstrain and GISAID databases and found that the L452R mutation is present in more than 400 SARS-CoV-2 genomes isolated from over 20 countries. More...
Emergence of a coronavirus SARS-CoV-2 E484K variant of interest in Arizona – VOI B.1.243.1
We sequenced the SARS-CoV-2 genome from 688 positive samples collected from December 28 2020 to March 13 2021 in Arizona, USA. More...
California coronavirus variant CAL.20C accounts for nearly half of COVID-19 cases in Southern California and about a 1/3 of cases in the state
The CAL.20C variant accounts for nearly half of COVID-19 cases in Southern California and about a third of cases in the state based on an analysis of viral genomes posted to a global database called GISAID. More...